- HOME
- Vienna 2024
- Lithuania 2023
- 16th CEEPC Prague 2022
- Proteome & Proteomics
- Proteomics Potentiality
- Precision Medicine & Cancer
- Proteomics and COVID-19
- Big Data & AI
- Spotlight Lithuania
- Humanity
- Meeting Reports & Tributes
- Country Profiles
- Proteomic Snippets
- Enabling Advances
- Spotlight Czech Republic
- Spotlight Poland
- 13th CEEPC - Ustron, Poland
- Spotlight Romania
- 12th CEEPC - Bucharest
- Spotlight Slovakia
- Spotlight Macedonia
- Sports Medicine
- Contacts & Copyrights
Power of Proteomics & 'Single Cell Proteomics' Research

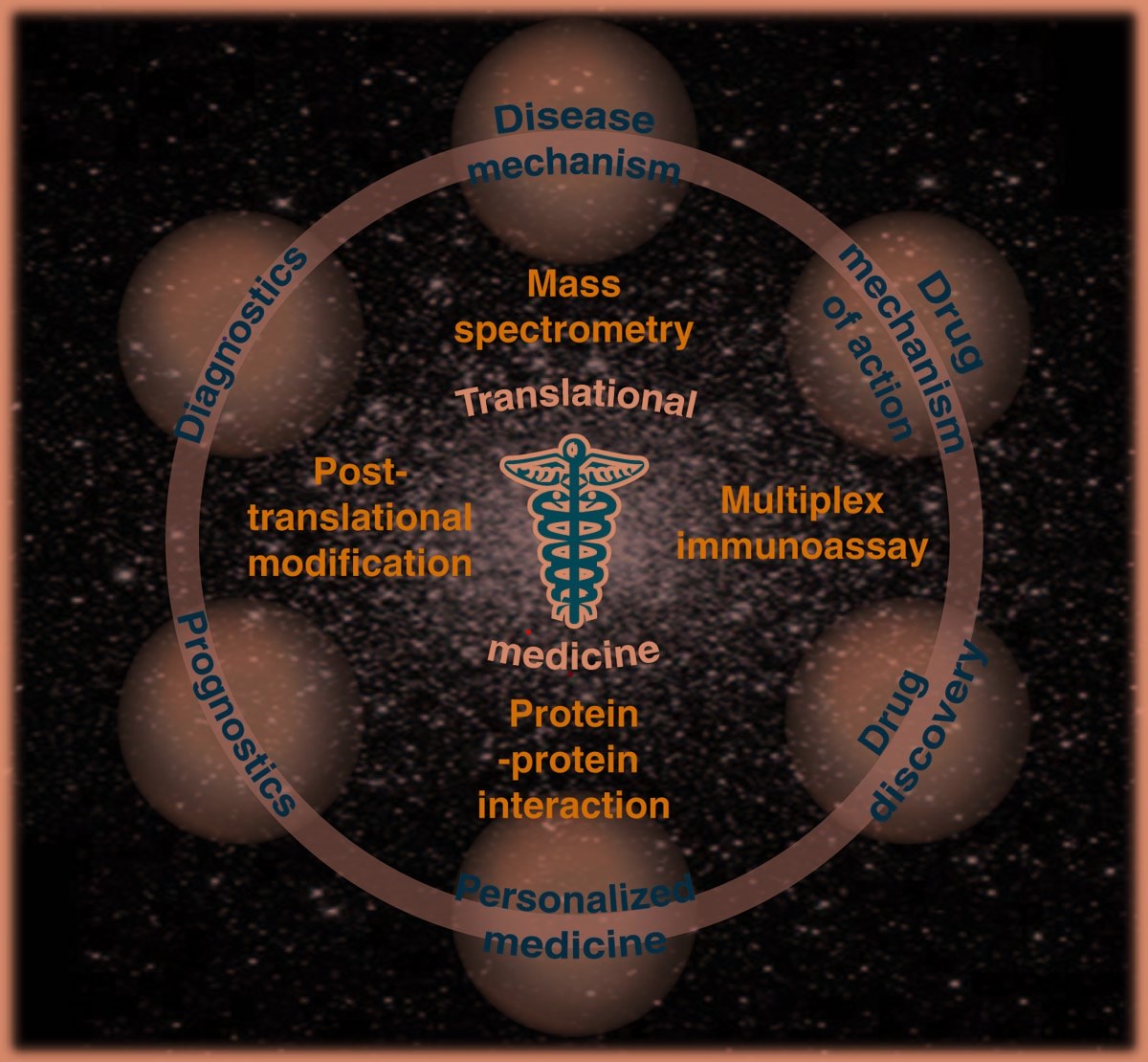

The goal of CEEPC is to promote the ‘state-of-the-art’ research in all types of Proteomic fields and it provides a scholarly forum where a whole plethora of proteomic disciplines are discussed, some examples of which are:
1. Single Cell Proteomics - highest quantitative accuracy and proteome depth interrogation with reliability.
2. Regenerative and Reparative Medicine: Stem cells, gene therapy; tissue engineering; cell based therapy; cell cultivation.
3. Disease Proteomics: Biopharmaceutical proteomics; diagnostics and prognostic marker studies; imaging; pharmacogenomics (personalized medicine); microarray technology; biomarkers, biopharmaceuticals discovery (CNS, cancer, cardiovascular, endocrine, immune); vaccines; antibodies; protein engineering.
4. Clinical Proteomics and Drug Discovery: in vivo and in vitro studies, cohort studies, clinical trials, disease diagnosis, data evaluation, drug design
5. Plant Proteomics and Environment: Transgenic plants and crops; bioremediation; microbial diversity; bio-monitoring; photosynthetic microorganisms, cyanobacteria and microalgae; genomics-assisted breeding.
6. Proteomics and tissue / protein engineering: connective tissue engineering, cell lines and stem cell technology, 'fed-batch culture' for biomarkers, bio-organs, tissue banking, industrial enzymes and proteins; bioprocess engineering and optimisation.
7. Proteomics and Bio-informatics: software, data analysis, protein databases, algorithms
8. Marine Proteomics: marine-based drug discovery & development
9. Additional Specialities: bio-safety; systems biology; bioethics; nanobioproteomics.
10. Quantitative Proteomics with digital intelligence - for biomarker research in health and disease
11. Stem cell Proteomics - for cell and gene therapy
12. AI driven Proteomics - in 'bundled Proteo-genomic-metabolomic' disease bimarker research.
Proteomics Prominence
Unlike the trend of many disciplines to join the '-omics' family by applying the suffix to their description, Proteomics together with Genomics and Transcriptomics had the luxury of prominence when researchers started to indulge in the exciting studies involving proteins and genes. Today Proteomics prominance stands tall because its twinning with other disciplines, and this gives it even a greater credibility as it facilitates the progression and direction of many disciplines sitting in 'silos' or often working in total isolation.
Disease research and Drug Discovery often focuses on human proteomics using multi-disciplinary approaches to untangle complex biological processes. Various studies on cellular proteins and protein interactions have evolved and being progressed due to advances in scientific knowledge and expansion of novel sophisticated technologies.
Proteomics Potentiality for diseases affecting mankind:
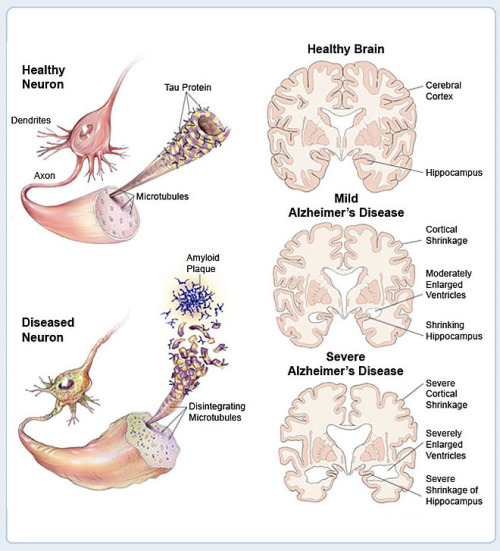
Neuro-Proteomics
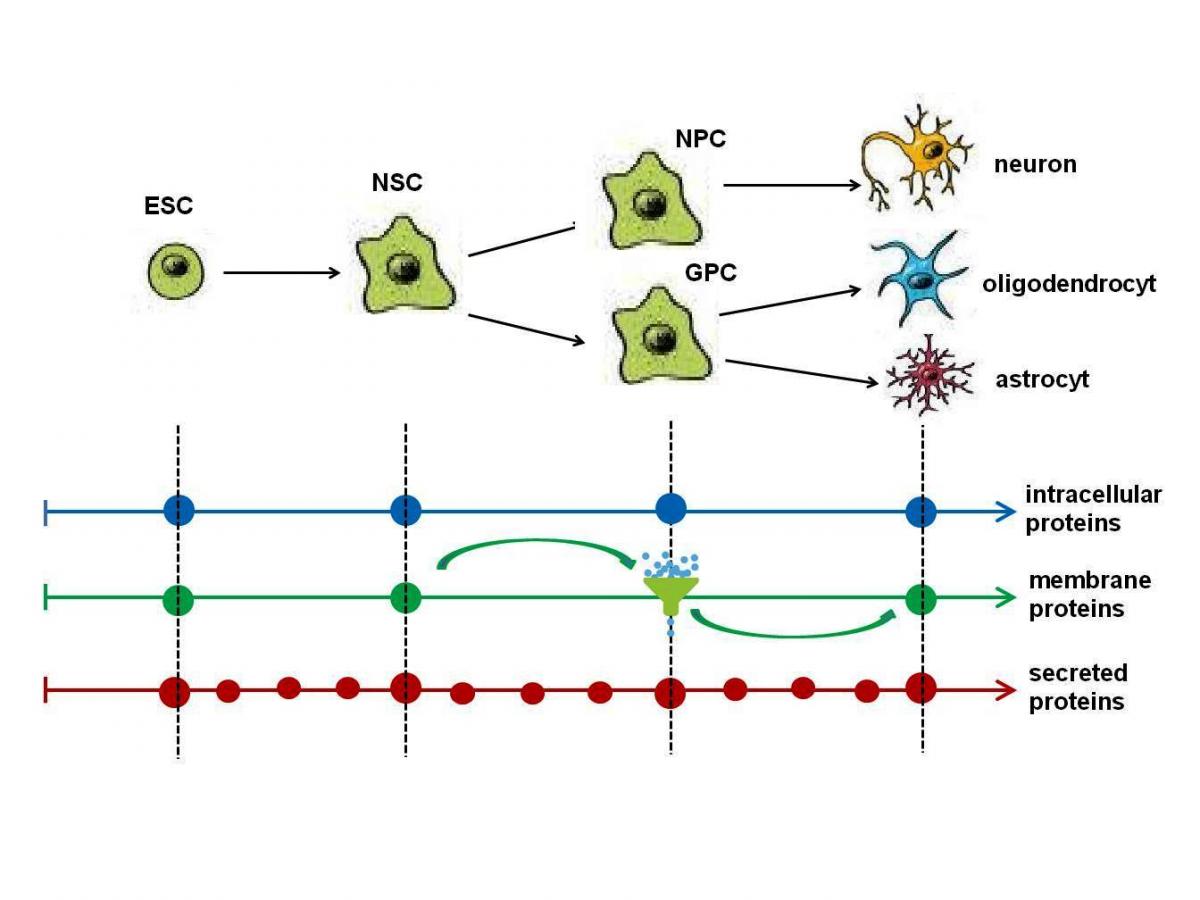
Neuroproteomics aims to study the expression, interaction and function of proteins in the nervous system using highly advanced technologies in combination with bioinformatics. Such approaches can help characterise neural networks in terms of their composition, organisation and dynamics related to physiological and behaviour processes. Importantly, this knowledge will contribute to the elucidation of a variety of neural disorders, including neurodegenerative disease where not only neurons but glial cells and many other subsets of the cells play a critical role in the functioning of the nervous system.
Proteo-Genomics
The completion of the human genome sequence more than a decade ago was a triumph for medical research; however, it did not provide all the answers it promised to tackle diseases and in fact, raised more questions. Today, extraordinary rate of advances in techniques for large scale automated genotyping is taking place where whole genome mRNA analysis and DNA sequencing has exceeded all expectations and has even exceeded the rate of developments in computing and data analysis. The idea of population based whole genome sequencing stands a reality. Connecting genotype to phenotype will require an ‘in-depth’ view of how the protein products of genes operate and interact. While large proteomic data may be collected from many different tissue types, focus is needed to consider tissue-specific proteins and post-translational modifications that affect protein-protein interactions, in addition to proteins produced from alternatively spliced transcripts. For disease research, Bioinformatics and data management are critical to providing a seamless, end-to-end sequencing solution, where efficiency, value of genomic analysis, and new methodologies and algorithms that link next-generation sequencing data to disease-relevant genetic markers is absolutely necessary for progress.
Proteo-Transcriptomics

It was believed that information transfer occurred from genome to transcriptome to proteome. Recent research has shown that it’s a two-way road—proteomics can be used to annotate the genome. The importance of this finding is that, using datasets, the understanding of the genome can be vastly improved with the help of algorithms that predict transcription and translation. Genomics and transcriptomics research can now hugely benefit from proteomics data and findings.
Metabolomics
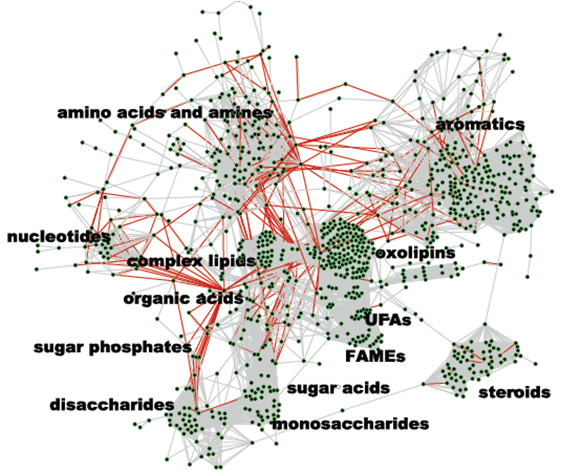
Metabolomics is a newly emerging field concerned with the comprehensive characterisation of the small molecule metabolites in biological systems. It can provide an overview of the metabolic status and overall biochemical events associated with a cellular or biological system. As such, it can accurately and comprehensively depict both the steady-state physiological state of a biological cell or organism and of their dynamic responses to genetic, abiotic and biotic environmental modulation. When Proteomic advances to date are utilised together with the rapidly expanding novel technologies in the understanding of the peptides and proteins, metabolomic expansion for the understanding of the role of small molecules in a dynamic system may significantly benefit.
Proteo-Cellomics
One of the greatest future technological challenges will be to apply Proteomic and Genomic techniques to single cells and to achieve a comparable level of sophistication of cellular analysis, to that we now have for working with DNA, RNA, and proteins.
Proteomics related Biomarker Discovery
Proteomics is the most applicable technology for implementing biomarker approaches in drug discovery given that virtually all existing drug targets are proteins. Proteomics and advancement in proteomic technologies has increased the chances of understanding the function of proteins that could have a role to play in the pathophysiology of diseases, thus enhancing the diagnostic or prognostic potential of such proteins as Biomarkers. Targeted proteomic quantification methodologies in understanding and unraveling various diseases including auto-immune and metabolic diseases, are now dependent on sophisticated technologies such as mass spectrometry approaches and extensive set of software tools for biomarker discovery.
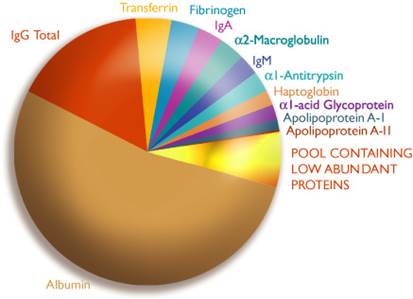 Given the complex nature and low abundance of many proteomic biomarkers, development of more sensitive methods and a massive integration of proteomic-based methodologies will continue to be needed if a biomarker(s) are to be discovered. Ideally, clinically useful biomarkers should be readily accessible in any body fluids such as follicular fluid, blood serum or urine, however, this is not the case. Hence novel and highly sensitive technologies still have a role to play in biomarker discovery.
Given the complex nature and low abundance of many proteomic biomarkers, development of more sensitive methods and a massive integration of proteomic-based methodologies will continue to be needed if a biomarker(s) are to be discovered. Ideally, clinically useful biomarkers should be readily accessible in any body fluids such as follicular fluid, blood serum or urine, however, this is not the case. Hence novel and highly sensitive technologies still have a role to play in biomarker discovery.
Proteomics related Bioinformatics

Proteomics related Bioinformatics has a major role to play in disease research where computational proteomics, next generation sequencing and sequence analysis and Computational systems for modelling biological processes are used. There is a great need for expanding the field of structural bioinformatics, developing important technologies and algorithms to solve specific problems in several bioinformatic topics. One such example is molecular bases of how macromolecular complexes and cell networks operate by analysing 'protein-protein' interaction networks with the help of novel computational techniques and methods, better data mining, data analysis and pattern recognition. All this could help understand the 'on-going' processes within the microenvironment in the cell. With reference to Genomics, next generation sequencing data analysis, protein and RNA structure understanding, could highlight protein function and proteomics comprehension. For complex disease scenarios, new approaches such as medical informatics and translational bioinformatics, computational systems biology, modelling and simulation and their application to the disease, may help researchers in their quest for answers.
Examples of 'power of softwares and Protein Databases' - identifying and linking biological systems for biomarker research
A
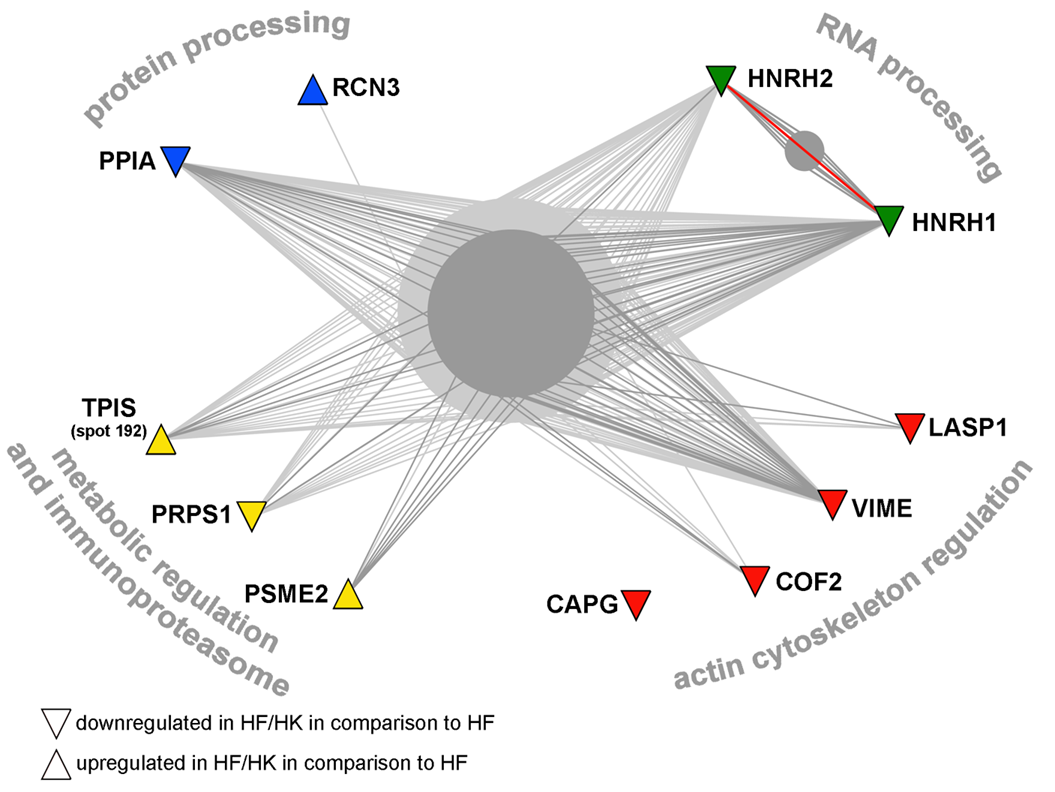 B
B
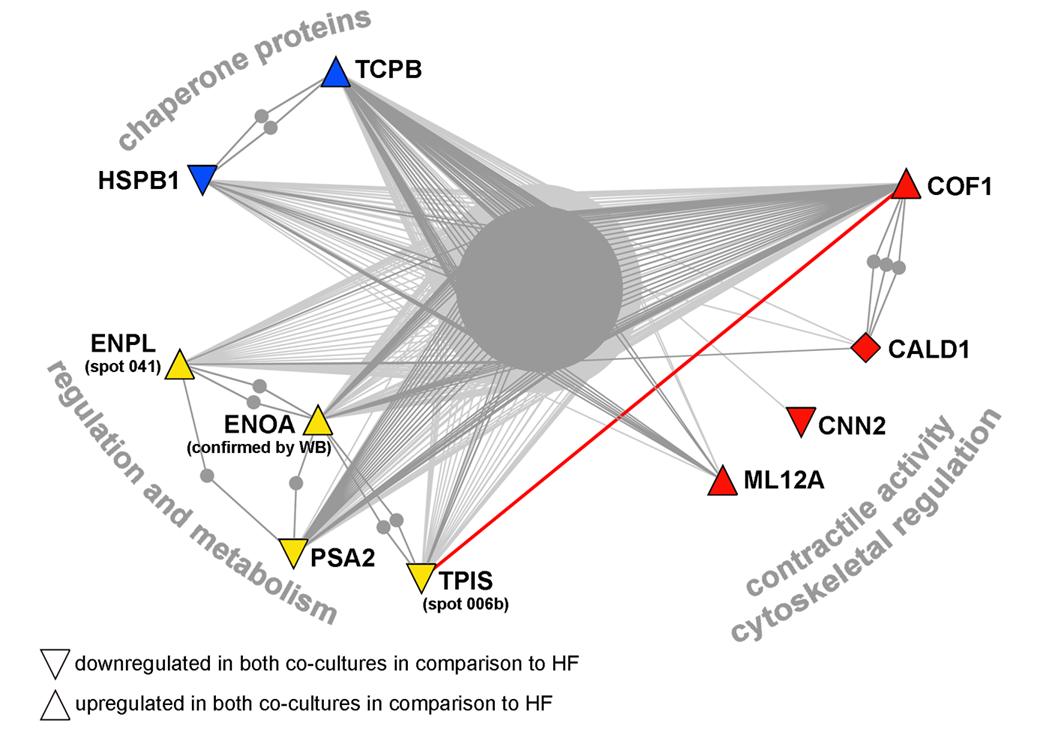
Example of protein interaction networks for the proteins in human fibroblasts influenced in co-culture models (A) or under the influence of normal keratinocytes only (B). A list of proteins was subjected to search in OPHID database and resulting graph files were processed in the NAViGaTOR software to visualise interaction networks. Different protein functional groups were highlighted by various colours, where red is for proteins associated with contractile activity and/or cytoskeleton, blue is for protein processing and chaperone proteins, green for RNA processing proteins, and yellow for proteins involved in regulation and metabolism. Thick red line highlights direct interaction between proteins identified in the study.
Regenerative Medicine for Degenerative Diseases
With aging population, today's challenges and diseases are acute. Below schematic attempts to highlight possible avenues to help resolve these issues
Reparative Medicine:
Use of Stem cells, gene therapy; tissue engineering; cell based therapy and cell cultivation can help tissue / organ repair. Many such attempt have provided a degree of success as shown below:
Treating damaged heart wall tissue by injecting stem cells:
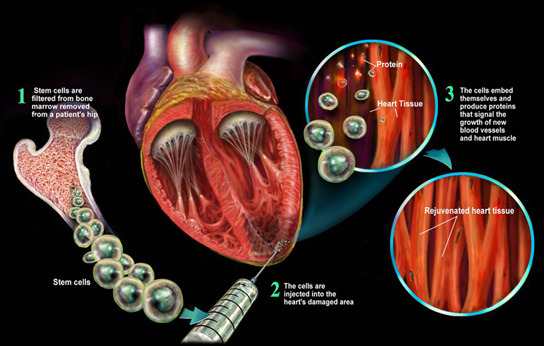
Stem Cell Proteomics
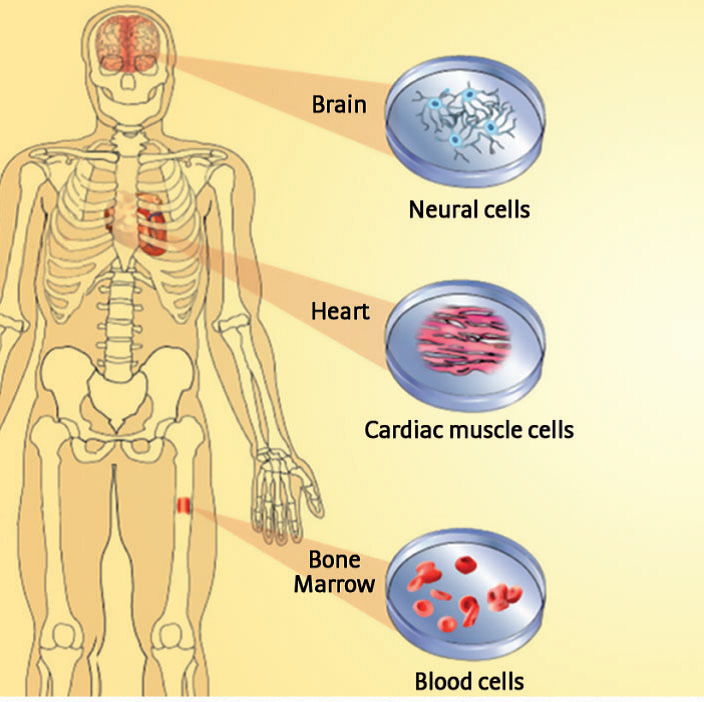
Advances in Stem Cell Research have highlighted the need for Proteomics where 'Stem Cell Proteomics' can play a major role in the understanding and application of this technology to diseases affecting mankind. Whilst the excitement remains high, the reality remains challenging ! - to read more about these challenges and the future of stem cell research, please click on the link below.
Special Issue: Focus on Stem Cell Proteomics October 2011 Volume 11, Issue 20, Pages 3943–4115

- Stem cell proteomics < Click link to read
“Identification of the human skeletal stem cell,” C.K.F. Chan et al., Cell, doi:10.1016/ j.cell.2018.07.029, 2018.
Human skeletal stem cells that become bone, cartilage, or stroma cells have been isolated from fetal and adult bones. This is the first time that skeletal stem cells, which had been observed in rodent models, have been identified in humans. The researchers were also able to derive the skeletal stem cells from human induced pluripotent stem cells, opening up new therapeutic possibilities for bone and cartilage related applications in diseases such as osteo and rheumatoid arthritis.
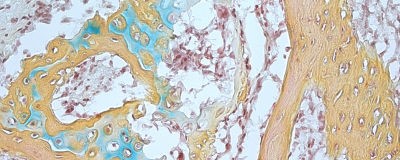
Image shows tissue derived from a single human skeletal stem cell. Bone is in yellow, blue indicates cartilage, and marrow is pictured in red.
Credit: CHAN AND LONGAKER et al.

Promoting regenerative and reparative medicine using Stem cells !
Stem Cell Research

Promoting exciting approches for benefit of mankind !
Gene deletion / Gene edition using CRISPR technology - exciting time, however, whole genome sequencing must be employed to look for all off-target effects in living models prior to any clinical approaches in humans are to be carried out.
